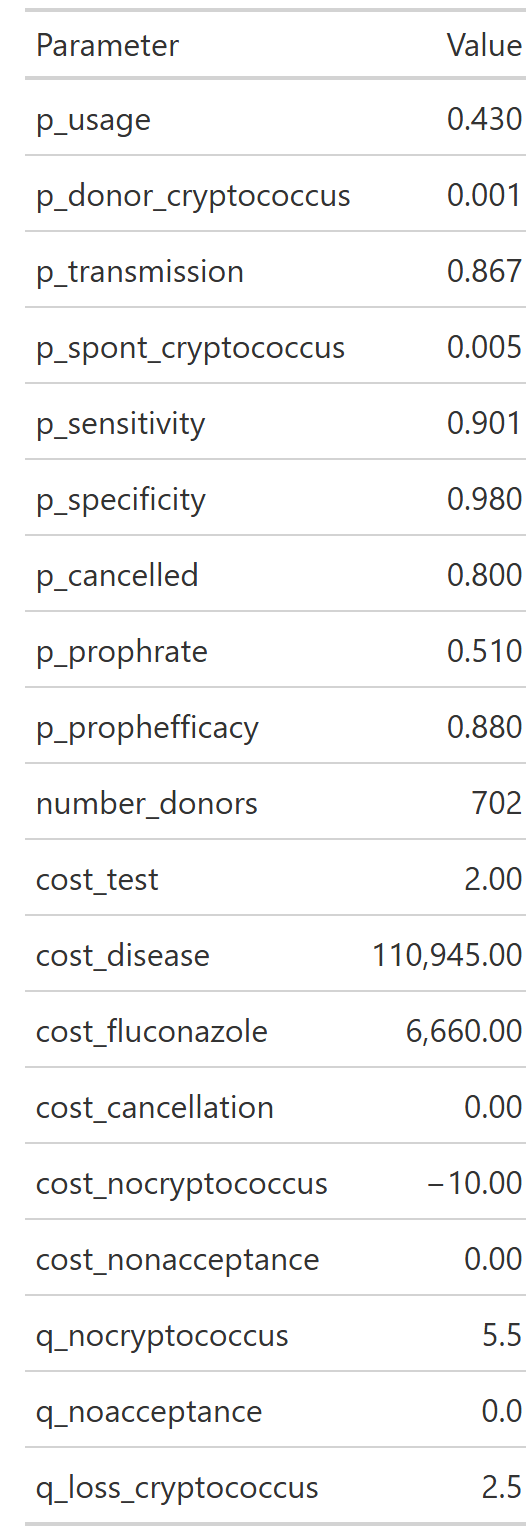
--- title: "Base case analysis" format: html --- ## What is a decision tree? ### Source code - [ `R/create_diagram_QC.R` ](https://github.com/VagishHemmige/Cryptococcus-donor-screening-CEA/blob/master/R/create_diagram_QC.R) - [ `R/setup.R` ](https://github.com/VagishHemmige/Cryptococcus-donor-screening-CEA/blob/master/R/setup.R) - [ `R/functions.R` ](https://github.com/VagishHemmige/Cryptococcus-donor-screening-CEA/blob/master/R/functions.R) ## Conceptual decision tree  {- Screening (e.g., CrAg or other donor testing strategy)- No screening (standard approach)- Donor truly has cryptococcal infection (rare)- Test is positive or negative (if screening)- Transmission occurs vs does not occur- Recipient develops disease vs not- Treatment works vs fails, etc.- Cost: testing costs, prophylaxis, hospitalization, antifungals, ICU care, etc.- QALYs: quality-adjusted survival for the recipient under that outcome## Model parameters ## Decision tree with parameters  ## Path table 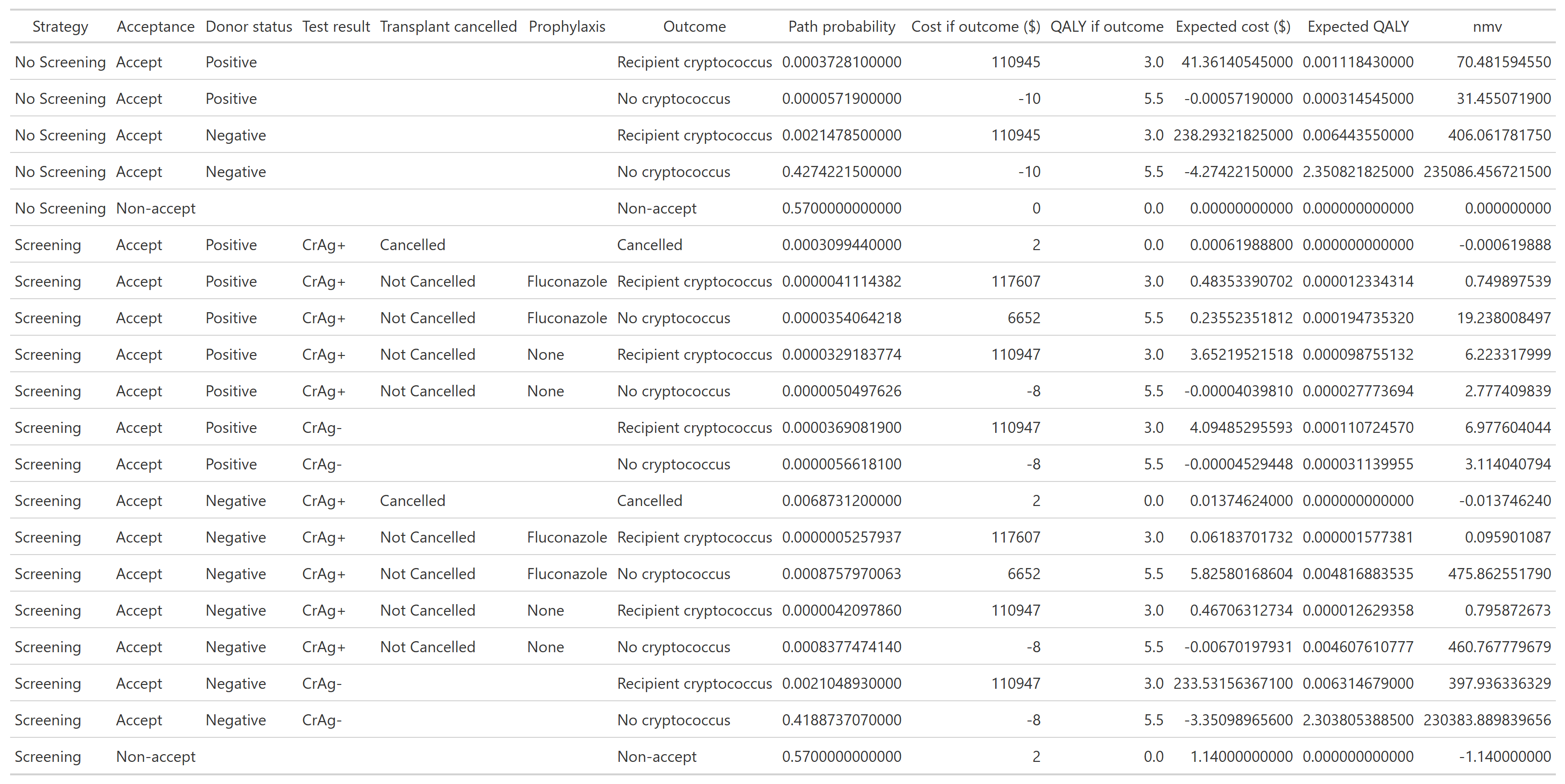 ## Summary table 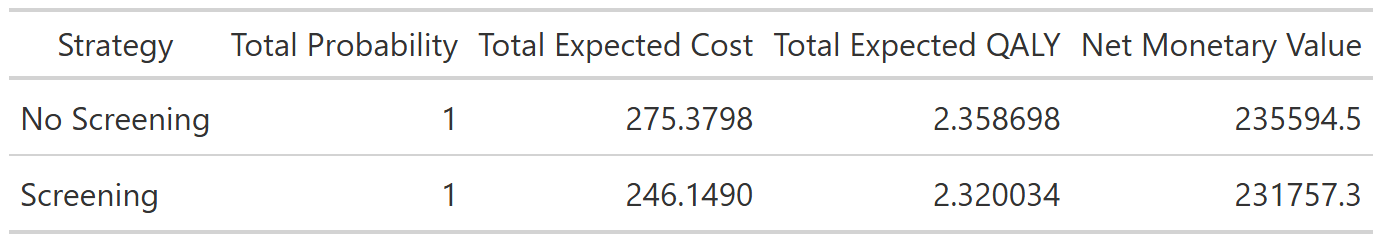 ## Source code [ `R/create_diagram_QC.R` ](https://github.com/VagishHemmige/Cryptococcus-donor-screening-CEA/blob/master/R/create_diagram_QC.R) that is used to perform this analysis. ```{r diagram, eval=FALSE} #Creates the tree diagram manually in R using grviz language #First, we source the helper functions for the analysis source("R/functions.R") #Run the tree diagram g_QC<-create_tree_diagram_QC() #Print plot g_QC$plot #Save the tree diagram as svg svg_QC <- export_svg(g_QC$plot) writeLines(svg_QC, "figures/crag_tree_QC.svg") #Annotate figure svg_doc <- read_xml(svg_QC) add_svg_annotation( svg_doc, c("Screening vs no screening:", glue("• ΔC = {scales::number(g_QC$cost_difference, accuracy = 0.01)}"), glue("• ΔQ = {formatC(g_QC$qaly_difference, format = 'f', digits = 4)}") ) ) write_xml(svg_doc, "figures/crag_tree_QC_annotated.svg") #Now, we look at the path table result_tibble_QC<-g_QC$path_table #Let's look at the tibble as a gt table crag_table_QC<-result_tibble_QC%>%gt()%>% cols_label( strategy = "Strategy", acceptance = "Acceptance", donor_dz_status = "Donor status", donor_test_result = "Test result", cancellation = "Transplant cancelled", proph = "Prophylaxis", outcome = "Outcome", probability = "Path probability", cost_total = "Cost if outcome ($)", cost_expected = "Expected cost ($)", qaly_total = "QALY if outcome", qaly_expected = "Expected QALY" )%>% tab_style( style = cell_text(align = "center"), locations = cells_column_labels(everything()) )%>% fmt_missing(everything(), missing_text = "") #Print the table crag_table_QC #Save table crag_table_QC%>% gtsave("figures/output_table_QC.png", vwidth = 2000, # try 1600–2400 vheight = 1200, expand = 10) crag_table_QC%>% gtsave("tables/output_table_QC.docx") #Let's create a table from the parameters parameter_table_QC<-g_QC$parameter_tibble%>% gt()%>% cols_label( parameter = "Parameter", value = "Value" )%>% fmt_number( columns = value, rows = parameter %in% c("number_donors"), decimals = 0 )%>% fmt_number( columns = value, rows = parameter %in% c("cost_test", "cost_disease", "cost_fluconazole", "cost_cancellation", "cost_cryptococcus", "cost_nonacceptance", "cost_nocryptococcus"), decimals = 2 )%>% fmt_number( columns = value, rows = parameter %in% c("q_nocryptococcus", "q_noacceptance", "q_loss_cryptococcus"), decimals = 1 ) parameter_table_QC parameter_table_QC%>% gtsave("figures/parameter_table_QC.png") parameter_table_QC%>% gtsave("tables/parameter_table_QC.docx") #Let's also look at the summary statistics for this table summary_tibble_QC<-result_tibble_QC%>% group_by(strategy)%>% summarize(total_probability=sum(probability), total_expected_cost=sum(cost_expected),total_expected_qaly=sum(qaly_expected)) summary_gt_QC<-summary_tibble_QC%>% gt()%>% cols_label( strategy = "Strategy", total_probability = "Total Probability", total_expected_cost = "Total Expected Cost", total_expected_qaly = "Total Expected QALY" )%>% tab_style( style = cell_text(align = "center"), locations = cells_column_labels(everything()) ) summary_gt_QC summary_gt_QC%>% gtsave("figures/summary_table_QC.png") summary_gt_QC%>% gtsave("tables/summary_table_QC.docx") ``` - [ `R/setup.R` ](https://github.com/VagishHemmige/Cryptococcus-donor-screening-CEA/blob/master/R/setup.R) ```{r setup, eval=FALSE} #This code performs setup and other tasks for the CEA analysis library(DiagrammeR) library(DiagrammeRsvg) library(glue) library(tibble) library(gt) library(tidyverse) library(ggtext) library(xml2) #Save simulation parameters #Expected values expected_value<-list() expected_value$p_usage<-0.43 expected_value$p_donor_cryptococcus<-0.001 expected_value$p_transmission<-0.867 expected_value$p_spont_cryptococcus<-0.005 expected_value$p_sensitivity<-0.901 expected_value$p_specificity<-0.98 expected_value$p_cancelled<-0.8 expected_value$p_prophrate<-0.51 expected_value$p_prophefficacy<-0.88 expected_value$number_donors<-702 expected_value$cost_test<-2 expected_value$cost_disease<-110945 expected_value$cost_fluconazole<-6660 expected_value$cost_cancellation<-0 expected_value$cost_nocryptococcus<--10 expected_value$cost_nonacceptance<-0 expected_value$q_nocryptococcus<-5.5 expected_value$q_noacceptance<-0 expected_value$q_loss_cryptococcus<-2.5 #Shape parameters shape_parameter<-list() shape_parameter$p_usage<-10 shape_parameter$p_donor_cryptococcus<-10 shape_parameter$p_transmission<-10 shape_parameter$p_spont_cryptococcus<-10 shape_parameter$p_sensitivity<-10 shape_parameter$p_specificity<-10 shape_parameter$p_cancelled<-10 shape_parameter$p_prophrate<-10 shape_parameter$p_prophefficacy<-10 shape_parameter$cost_test<-4 shape_parameter$cost_disease<-4 shape_parameter$cost_fluconazole<-4 shape_parameter$cost_cancellation<-NA shape_parameter$cost_nocryptococcus<-NA shape_parameter$cost_nonacceptance<-NA shape_parameter$q_nocryptococcus<-4 shape_parameter$q_noacceptance<-NA shape_parameter$q_loss_cryptococcus<-4 # willingness-to-pay threshold wtp <- 100000 #Set seed for reproducibility set.seed(42) #Define number of simulations in PSA nsim<-10000 ``` - [ `R/functions.R` ](https://github.com/VagishHemmige/Cryptococcus-donor-screening-CEA/blob/master/R/function.R) ```{r functions, eval=FALSE} #Functions for CEA #Function for the calculations needed for PSA. calculate_cost_QALY_QC<-function(p_usage=expected_value$p_usage, p_donor_cryptococcus=expected_value$p_donor_cryptococcus, p_transmission=expected_value$p_transmission, p_spont_cryptococcus=expected_value$p_spont_cryptococcus, p_sensitivity=expected_value$p_sensitivity, p_specificity=expected_value$p_specificity, p_cancelled=expected_value$p_cancelled, p_prophrate=expected_value$p_prophrate, p_prophefficacy=expected_value$p_prophefficacy, number_donors=expected_value$number_donors, cost_test=expected_value$cost_test, cost_disease=expected_value$cost_disease, cost_fluconazole=expected_value$cost_fluconazole, cost_cancellation=expected_value$cost_cancellation, cost_nocryptococcus=expected_value$cost_nocryptococcus, cost_nonacceptance=expected_value$cost_nonacceptance, q_nocryptococcus=expected_value$q_nocryptococcus, q_noacceptance=expected_value$q_noacceptance, q_loss_cryptococcus=expected_value$q_loss_cryptococcus ) { #Define parameters which are derived from the input parameters p_nonusage<-1-p_usage p_donor_nocryptococcus<-1-p_donor_cryptococcus p_nontransmission<-1-p_transmission p_nospont_cryptococcus<-1-p_spont_cryptococcus p_falsenegative<-1-p_sensitivity p_falsepositive<-1-p_specificity p_nocancelled<-1-p_cancelled p_noprophrate<-1-p_prophrate p_noprophefficacy<-1-p_prophefficacy p_breakthrough_donorpos<-(1-p_prophefficacy)*p_transmission p_nobreakthrough_donorpos<-1-p_breakthrough_donorpos p_breakthrough_donorneg<-(1-p_prophefficacy)*p_spont_cryptococcus p_nobreakthrough_donorneg<-1-p_breakthrough_donorneg q_cryptococcus<-q_nocryptococcus-q_loss_cryptococcus #Define tibble of outcomes path_table<-tribble( ~strategy, ~acceptance, ~donor_dz_status, ~donor_test_result, ~cancellation, ~proph, ~outcome, ~probability, ~cost_total, ~qaly_total, "No Screening","Accept","Positive",NA,NA,NA,"Recipient cryptococcus",p_usage*p_donor_cryptococcus*p_transmission,cost_disease,q_cryptococcus, "No Screening","Accept","Positive",NA,NA,NA,"No cryptococcus",p_usage*p_donor_cryptococcus*p_nontransmission,cost_nocryptococcus,q_nocryptococcus, "No Screening","Accept","Negative",NA,NA,NA,"Recipient cryptococcus",p_usage*p_donor_nocryptococcus*p_spont_cryptococcus,cost_disease,q_cryptococcus, "No Screening","Accept","Negative",NA,NA,NA,"No cryptococcus",p_usage*p_donor_nocryptococcus*p_nospont_cryptococcus,cost_nocryptococcus,q_nocryptococcus, "No Screening","Non-accept",NA,NA,NA,NA,"Non-accept",p_nonusage,cost_nonacceptance,q_noacceptance, "Screening","Accept","Positive","CrAg+","Cancelled",NA,"Cancelled",p_usage*p_donor_cryptococcus*p_sensitivity*p_cancelled,cost_cancellation+cost_test,q_noacceptance, "Screening","Accept","Positive","CrAg+","Not Cancelled","Fluconazole","Recipient cryptococcus",p_usage*p_donor_cryptococcus*p_sensitivity*p_nocancelled*p_prophrate*p_breakthrough_donorpos,cost_test+cost_disease+cost_fluconazole,q_cryptococcus, "Screening","Accept","Positive","CrAg+","Not Cancelled","Fluconazole","No cryptococcus",p_usage*p_donor_cryptococcus*p_sensitivity*p_nocancelled*p_prophrate*p_nobreakthrough_donorpos,cost_test+cost_nocryptococcus+cost_fluconazole,q_nocryptococcus, "Screening","Accept","Positive","CrAg+","Not Cancelled","None","Recipient cryptococcus",p_usage*p_donor_cryptococcus*p_sensitivity*p_nocancelled*p_noprophrate*p_transmission,cost_test+cost_disease,q_cryptococcus, "Screening","Accept","Positive","CrAg+","Not Cancelled","None","No cryptococcus",p_usage*p_donor_cryptococcus*p_sensitivity*p_nocancelled*p_noprophrate*p_nontransmission,cost_test+cost_nocryptococcus,q_nocryptococcus, "Screening","Accept","Positive","CrAg-",NA,NA,"Recipient cryptococcus",p_usage*p_donor_cryptococcus*p_falsenegative*p_transmission,cost_test+cost_disease,q_cryptococcus, "Screening","Accept","Positive","CrAg-",NA,NA,"No cryptococcus",p_usage*p_donor_cryptococcus*p_falsenegative*p_nontransmission,cost_test+cost_nocryptococcus,q_nocryptococcus, "Screening","Accept","Negative","CrAg+","Cancelled",NA,"Cancelled",p_usage*p_donor_nocryptococcus*p_falsepositive*p_cancelled,cost_cancellation+cost_test,q_noacceptance, "Screening","Accept","Negative","CrAg+","Not Cancelled","Fluconazole","Recipient cryptococcus",p_usage*p_donor_nocryptococcus*p_falsepositive*p_nocancelled*p_prophrate*p_breakthrough_donorneg,cost_test+cost_disease+cost_fluconazole,q_cryptococcus, "Screening","Accept","Negative","CrAg+","Not Cancelled","Fluconazole","No cryptococcus",p_usage*p_donor_nocryptococcus*p_falsepositive*p_nocancelled*p_prophrate*p_nobreakthrough_donorneg,cost_test+cost_nocryptococcus+cost_fluconazole,q_nocryptococcus, "Screening","Accept","Negative","CrAg+","Not Cancelled","None","Recipient cryptococcus",p_usage*p_donor_nocryptococcus*p_falsepositive*p_nocancelled*p_noprophrate*p_spont_cryptococcus,cost_test+cost_disease,q_cryptococcus, "Screening","Accept","Negative","CrAg+","Not Cancelled","None","No cryptococcus",p_usage*p_donor_nocryptococcus*p_falsepositive*p_nocancelled*p_noprophrate*p_nospont_cryptococcus,cost_test+cost_nocryptococcus,q_nocryptococcus, "Screening","Accept","Negative","CrAg-",NA,NA,"Recipient cryptococcus",p_usage*p_donor_nocryptococcus*p_specificity*p_spont_cryptococcus,cost_test+cost_disease,q_cryptococcus, "Screening","Accept","Negative","CrAg-",NA,NA,"No cryptococcus",p_usage*p_donor_nocryptococcus*p_specificity*p_nospont_cryptococcus,cost_test+cost_nocryptococcus,q_nocryptococcus, "Screening","Non-accept",NA,NA,NA,NA,"Non-accept",p_nonusage,cost_nonacceptance+cost_test,q_noacceptance, )%>% mutate(cost_expected=probability*cost_total, qaly_expected=probability*qaly_total) #Summary object is created via summarize and then converted into long format summary_tibble<-path_table%>% group_by(strategy)%>% summarize(total_expected_cost=sum(cost_expected), total_expected_qaly=sum(qaly_expected))%>% mutate( strategy = recode( strategy, "No Screening" = "ns", "Screening" = "s" ) ) %>% tidyr::pivot_wider( names_from = strategy, values_from = c( total_expected_cost, total_expected_qaly ), names_sep = "_" ) #Final return return(summary_tibble) } #Function that creates tree diagram. It returns the diagram created via the GraphViz language, a table of input parameters #and the output results create_tree_diagram_QC<-function(p_usage=expected_value$p_usage, p_donor_cryptococcus=expected_value$p_donor_cryptococcus, p_transmission=expected_value$p_transmission, p_spont_cryptococcus=expected_value$p_spont_cryptococcus, p_sensitivity=expected_value$p_sensitivity, p_specificity=expected_value$p_specificity, p_cancelled=expected_value$p_cancelled, p_prophrate=expected_value$p_prophrate, p_prophefficacy=expected_value$p_prophefficacy, number_donors=expected_value$number_donors, cost_test=expected_value$cost_test, cost_disease=expected_value$cost_disease, cost_fluconazole=expected_value$cost_fluconazole, cost_cancellation=expected_value$cost_cancellation, cost_nocryptococcus=expected_value$cost_nocryptococcus, cost_nonacceptance=expected_value$cost_nonacceptance, q_nocryptococcus=expected_value$q_nocryptococcus, q_noacceptance=expected_value$q_noacceptance, q_loss_cryptococcus=expected_value$q_loss_cryptococcus, circle_diameter=1.4, box_width=1.7, box_height=1, diamond_width=3, diamond_height=1.5 ) { #Define parameters that are derived from parameters passed to the function p_nonusage<-1-p_usage p_donor_nocryptococcus<-1-p_donor_cryptococcus p_nontransmission<-1-p_transmission p_nospont_cryptococcus<-1-p_spont_cryptococcus p_falsenegative<-1-p_sensitivity p_falsepositive<-1-p_specificity p_nocancelled<-1-p_cancelled p_noprophrate<-1-p_prophrate p_noprophefficacy<-1-p_prophefficacy p_breakthrough_donorpos<-(1-p_prophefficacy)*p_transmission p_nobreakthrough_donorpos<-1-p_breakthrough_donorpos p_breakthrough_donorneg<-(1-p_prophefficacy)*p_spont_cryptococcus p_nobreakthrough_donorneg<-1-p_breakthrough_donorneg q_cryptococcus<-q_nocryptococcus-q_loss_cryptococcus #Define return list return_list<-list() #Define tibble of outcomes return_list$path_table<-tribble( ~strategy, ~acceptance, ~donor_dz_status, ~donor_test_result, ~cancellation, ~proph, ~outcome, ~probability, ~cost_total, ~qaly_total, "No Screening","Accept","Positive",NA,NA,NA,"Recipient cryptococcus",p_usage*p_donor_cryptococcus*p_transmission,cost_disease,q_cryptococcus, "No Screening","Accept","Positive",NA,NA,NA,"No cryptococcus",p_usage*p_donor_cryptococcus*p_nontransmission,cost_nocryptococcus,q_nocryptococcus, "No Screening","Accept","Negative",NA,NA,NA,"Recipient cryptococcus",p_usage*p_donor_nocryptococcus*p_spont_cryptococcus,cost_disease,q_cryptococcus, "No Screening","Accept","Negative",NA,NA,NA,"No cryptococcus",p_usage*p_donor_nocryptococcus*p_nospont_cryptococcus,cost_nocryptococcus,q_nocryptococcus, "No Screening","Non-accept",NA,NA,NA,NA,"Non-accept",p_nonusage,cost_nonacceptance,q_noacceptance, "Screening","Accept","Positive","CrAg+","Cancelled",NA,"Cancelled",p_usage*p_donor_cryptococcus*p_sensitivity*p_cancelled,cost_cancellation+cost_test,q_noacceptance, "Screening","Accept","Positive","CrAg+","Not Cancelled","Fluconazole","Recipient cryptococcus",p_usage*p_donor_cryptococcus*p_sensitivity*p_nocancelled*p_prophrate*p_breakthrough_donorpos,cost_test+cost_disease+cost_fluconazole,q_cryptococcus, "Screening","Accept","Positive","CrAg+","Not Cancelled","Fluconazole","No cryptococcus",p_usage*p_donor_cryptococcus*p_sensitivity*p_nocancelled*p_prophrate*p_nobreakthrough_donorpos,cost_test+cost_nocryptococcus+cost_fluconazole,q_nocryptococcus, "Screening","Accept","Positive","CrAg+","Not Cancelled","None","Recipient cryptococcus",p_usage*p_donor_cryptococcus*p_sensitivity*p_nocancelled*p_noprophrate*p_transmission,cost_test+cost_disease,q_cryptococcus, "Screening","Accept","Positive","CrAg+","Not Cancelled","None","No cryptococcus",p_usage*p_donor_cryptococcus*p_sensitivity*p_nocancelled*p_noprophrate*p_nontransmission,cost_test+cost_nocryptococcus,q_nocryptococcus, "Screening","Accept","Positive","CrAg-",NA,NA,"Recipient cryptococcus",p_usage*p_donor_cryptococcus*p_falsenegative*p_transmission,cost_test+cost_disease,q_cryptococcus, "Screening","Accept","Positive","CrAg-",NA,NA,"No cryptococcus",p_usage*p_donor_cryptococcus*p_falsenegative*p_nontransmission,cost_test+cost_nocryptococcus,q_nocryptococcus, "Screening","Accept","Negative","CrAg+","Cancelled",NA,"Cancelled",p_usage*p_donor_nocryptococcus*p_falsepositive*p_cancelled,cost_cancellation+cost_test,q_noacceptance, "Screening","Accept","Negative","CrAg+","Not Cancelled","Fluconazole","Recipient cryptococcus",p_usage*p_donor_nocryptococcus*p_falsepositive*p_nocancelled*p_prophrate*p_breakthrough_donorneg,cost_test+cost_disease+cost_fluconazole,q_cryptococcus, "Screening","Accept","Negative","CrAg+","Not Cancelled","Fluconazole","No cryptococcus",p_usage*p_donor_nocryptococcus*p_falsepositive*p_nocancelled*p_prophrate*p_nobreakthrough_donorneg,cost_test+cost_nocryptococcus+cost_fluconazole,q_nocryptococcus, "Screening","Accept","Negative","CrAg+","Not Cancelled","None","Recipient cryptococcus",p_usage*p_donor_nocryptococcus*p_falsepositive*p_nocancelled*p_noprophrate*p_spont_cryptococcus,cost_test+cost_disease,q_cryptococcus, "Screening","Accept","Negative","CrAg+","Not Cancelled","None","No cryptococcus",p_usage*p_donor_nocryptococcus*p_falsepositive*p_nocancelled*p_noprophrate*p_nospont_cryptococcus,cost_test+cost_nocryptococcus,q_nocryptococcus, "Screening","Accept","Negative","CrAg-",NA,NA,"Recipient cryptococcus",p_usage*p_donor_nocryptococcus*p_specificity*p_spont_cryptococcus,cost_test+cost_disease,q_cryptococcus, "Screening","Accept","Negative","CrAg-",NA,NA,"No cryptococcus",p_usage*p_donor_nocryptococcus*p_specificity*p_nospont_cryptococcus,cost_test+cost_nocryptococcus,q_nocryptococcus, "Screening","Non-accept",NA,NA,NA,NA,"Non-accept",p_nonusage,cost_nonacceptance+cost_test,q_noacceptance, )%>% mutate(cost_expected=probability*cost_total, qaly_expected=probability*qaly_total) #Summary object ofr return_list$summary_tibble<-return_list$path_table%>% group_by(strategy)%>% summarize(total_probability=sum(probability), total_expected_cost=sum(cost_expected), total_expected_qaly=sum(qaly_expected)) #Create parameter table to return return_list$parameter_tibble<-tribble( ~parameter, ~value, "p_usage", p_usage, "p_donor_cryptococcus",p_donor_cryptococcus, "p_transmission",p_transmission, "p_spont_cryptococcus",p_spont_cryptococcus, "p_sensitivity",p_sensitivity, "p_specificity",p_specificity, "p_cancelled",p_cancelled, "p_prophrate",p_prophrate, "p_prophefficacy",p_prophefficacy, "number_donors",number_donors, "cost_test",cost_test, "cost_disease",cost_disease, "cost_fluconazole",cost_fluconazole, "cost_cancellation",cost_cancellation, "cost_nocryptococcus",cost_nocryptococcus, "cost_nonacceptance",cost_nonacceptance, "q_nocryptococcus",q_nocryptococcus, "q_noacceptance",q_noacceptance, "q_loss_cryptococcus",q_loss_cryptococcus) return_list$cost_difference<-return_list$summary_tibble[[2,3]]-return_list$summary_tibble[[1,3]] return_list$qaly_difference<-return_list$summary_tibble[[2,4]]-return_list$summary_tibble[[1,4]] #Text to define tree diagram for grviz grviz_text<-glue(" digraph crag {{ graph [rankdir=LR] node [fontname=Helvetica, fontsize=14.5] edge [fontname=Helvetica, fontsize=15] # ---- Nodes ---- start [shape=box, label='Potential\ndonors\nN = {number_donors}', fixedsize=TRUE, width={box_width}, height={box_height}] no_screen [shape=box, fillcolor=palegreen, style=filled, label='No CrAg\nscreening\nΔC = $0', fixedsize=TRUE, width={box_width}, height={box_height}] screen [shape=box, fillcolor=palegreen, style=filled, label='CrAg\nscreening\nΔC = ${cost_test}', fixedsize=TRUE, width={box_width}, height={box_height}] used_ns [shape=circle, fillcolor=lightyellow, style=filled, label='Organs\nused', fixedsize=TRUE, width={circle_diameter}, height={circle_diameter}] notused_ns [shape=diamond, fillcolor=lightblue, style=filled, label='Organs\nnot used\nΔC = ${cost_nonacceptance}\nΔQ = {q_noacceptance}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] used_s [shape=circle, fillcolor=lightyellow, style=filled, label='Organs\nused', fixedsize=TRUE, width={circle_diameter}, height={circle_diameter}] notused_s [shape=diamond, fillcolor=lightblue, style=filled, label='Organs\nnot used\nΔC = ${cost_nonacceptance}\nΔQ = {q_noacceptance}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] donor_pos_ns [shape=circle, fillcolor=lightyellow, style=filled, label='Donor\nwith\ncryptococcus', fixedsize=TRUE, width={circle_diameter}, height={circle_diameter}] donor_neg_ns [shape=circle, fillcolor=lightyellow, style=filled, label='Donor\nwithout\ncryptococcus', fixedsize=TRUE, width={circle_diameter}, height={circle_diameter}] donor_pos_s [shape=circle, fillcolor=lightyellow, style=filled, label='Donor\nwith\ncryptococcus', fixedsize=TRUE, width={circle_diameter}, height={circle_diameter}] donor_neg_s [shape=circle, fillcolor=lightyellow, style=filled, label='Donor\nwithout\ncryptococcus', fixedsize=TRUE, width={circle_diameter}, height={circle_diameter}] inf_yes_agpos_ns [shape=diamond, fillcolor=mistyrose, style=filled, label='Recipient\ncryptococcosis\nΔC = ${cost_disease}\nΔQ = {q_cryptococcus}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] inf_yes_agneg_ns [shape=diamond, fillcolor=mistyrose, style=filled, label='Recipient\ncryptococcosis\nΔC = ${cost_disease}\nΔQ = {q_cryptococcus}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] inf_no_agpos_ns [shape=diamond, label='No\ncryptococcus\nΔC = ${cost_nocryptococcus}\nΔQ = {q_nocryptococcus}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] inf_no_agneg_ns [shape=diamond, label='No\ncryptococcus\nΔC = ${cost_nocryptococcus}\nΔQ = {q_nocryptococcus}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] ag_truepos_s [shape=circle, fillcolor=lightyellow, style=filled, label='Donor\nCrAg+', fixedsize=TRUE, width={circle_diameter}, height={circle_diameter}] ag_falseneg_s [shape=circle, fillcolor=lightyellow, style=filled, label='Donor\nCrAg-', fixedsize=TRUE, width={circle_diameter}, height={circle_diameter}] ag_falsepos_s [shape=circle, fillcolor=lightyellow, style=filled, label='Donor\nCrAg+', fixedsize=TRUE, width={circle_diameter}, height={circle_diameter}] ag_trueneg_s [shape=circle, fillcolor=lightyellow, style=filled, label='Donor\nCrAg-', fixedsize=TRUE, width={circle_diameter}, height={circle_diameter}] transplant_cancelled_ag_truepos_s [shape=diamond, fillcolor=lightblue, style=filled, label='Transplant\ncancelled\nΔC = ${cost_cancellation}\nΔQ = {q_noacceptance}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] transplant_notcancelled_ag_truepos_s [shape=circle, fillcolor=lightyellow, style=filled, label='Transplant\nnot\ncancelled', fixedsize=TRUE, width={circle_diameter}, height={circle_diameter}] transplant_cancelled_ag_falsepos_s [shape=diamond, fillcolor=lightblue, style=filled, label='Transplant\ncancelled\nΔC = ${cost_cancellation}\nΔQ = {q_noacceptance}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] transplant_notcancelled_ag_falsepos_s [shape=circle, fillcolor=lightyellow, style=filled, label='Transplant\nnot\ncancelled', fixedsize=TRUE, width={circle_diameter}, height={circle_diameter}] proph_yes_transplant_notcancelled_ag_truepos_s [shape=circle, fillcolor=lightyellow, style=filled, label='Fluconazole\nprophylaxis\nΔC = ${cost_fluconazole}', fixedsize=TRUE, width={circle_diameter}, height={circle_diameter}] proph_no_transplant_notcancelled_ag_truepos_s [shape=circle, fillcolor=lightyellow, style=filled, label='No\nprophylaxis', fixedsize=TRUE, width={circle_diameter}, height={circle_diameter}] proph_yes_transplant_notcancelled_ag_falsepos_s [shape=circle, fillcolor=lightyellow, style=filled, label='Fluconazole\nprophylaxis\nΔC = ${cost_fluconazole}', fixedsize=TRUE, width={circle_diameter}, height={circle_diameter}] proph_no_transplant_notcancelled_ag_falsepos_s [shape=circle, fillcolor=lightyellow, style=filled, label='No\nprophylaxis', fixedsize=TRUE, width={circle_diameter}, height={circle_diameter}] inf_y_proph_yes_transplant_notcancelled_ag_truepos_s [shape=diamond, fillcolor=mistyrose, style=filled, label='Recipient\ncryptococcosis\nΔC = ${cost_disease}\nΔQ = {q_cryptococcus}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] inf_n_proph_yes_transplant_notcancelled_ag_truepos_s [shape=diamond, label='No\ncryptococcosis\nΔC = ${cost_nocryptococcus}\nΔQ = {q_nocryptococcus}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] inf_y_proph_no_transplant_notcancelled_ag_truepos_s [shape=diamond, fillcolor=mistyrose, style=filled, label='Recipient\ncryptococcosis\nΔC = ${cost_disease}\nΔQ = {q_cryptococcus}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] inf_n_proph_no_transplant_notcancelled_ag_truepos_s [shape=diamond, label='No\ncryptococcosis\nΔC = ${cost_nocryptococcus}\nΔQ = {q_nocryptococcus}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] inf_y_proph_yes_transplant_notcancelled_ag_falsepos_s [shape=diamond, fillcolor=mistyrose, style=filled, label='Recipient\ncryptococcosis\nΔC = ${cost_disease}\nΔQ = {q_cryptococcus}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] inf_n_proph_yes_transplant_notcancelled_ag_falsepos_s [shape=diamond, label='No\ncryptococcus\nΔC = ${cost_nocryptococcus}\nΔQ = {q_nocryptococcus}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] inf_y_proph_no_transplant_notcancelled_ag_falsepos_s [shape=diamond, fillcolor=mistyrose, style=filled, label='Recipient\ncryptococcosis\nΔC = ${cost_disease}\nΔQ = {q_cryptococcus}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] inf_n_proph_no_transplant_notcancelled_ag_falsepos_s [shape=diamond, label='No\ncryptococcus\nΔC = ${cost_nocryptococcus}\nΔQ = {q_nocryptococcus}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] inf_y_ag_falseneg_s [shape=diamond, fillcolor=mistyrose, style=filled, label='Recipient\ncryptococcosis\nΔC = ${cost_disease}\nΔQ = {q_cryptococcus}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] inf_n_ag_falseneg_s [shape=diamond, label='No\ncryptococcus\nΔC = ${cost_nocryptococcus}\nΔQ = {q_nocryptococcus}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] inf_y_ag_trueneg_s [shape=diamond, fillcolor=mistyrose, style=filled, label='Recipient\ncryptococcosis\nΔC = ${cost_disease}\nΔQ = {q_cryptococcus}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] inf_n_ag_trueneg_s [shape=diamond, label='No\ncryptococcus\nΔC = ${cost_nocryptococcus}\nΔQ = {q_nocryptococcus}', fixedsize=TRUE, width={diamond_width}, height={diamond_height}] # ---- Structure ---- start -> no_screen start -> screen no_screen -> used_ns [label='p = {p_usage}'] no_screen -> notused_ns [label='p = {p_nonusage}'] used_ns -> donor_pos_ns [label='p = {p_donor_cryptococcus}'] used_ns -> donor_neg_ns [label='p = {p_donor_nocryptococcus}'] donor_pos_ns -> inf_yes_agpos_ns [label='p = {p_transmission}'] donor_pos_ns -> inf_no_agpos_ns [label='p = {p_nontransmission}'] donor_neg_ns -> inf_no_agneg_ns [label='p = {p_nospont_cryptococcus}'] donor_neg_ns -> inf_yes_agneg_ns [label='p = {p_spont_cryptococcus}'] screen -> used_s [label='p = {p_usage}'] screen -> notused_s [label='p = {p_nonusage}'] used_s -> donor_pos_s [label='p = {p_donor_cryptococcus}'] used_s -> donor_neg_s [label='p = {p_donor_nocryptococcus}'] donor_pos_s -> ag_truepos_s [label='p = {p_sensitivity}'] donor_pos_s -> ag_falseneg_s [label='p = {p_falsenegative}'] ag_truepos_s -> transplant_cancelled_ag_truepos_s [label='p = {p_cancelled}'] ag_truepos_s -> transplant_notcancelled_ag_truepos_s [label='p = {p_nocancelled}'] transplant_notcancelled_ag_truepos_s-> proph_yes_transplant_notcancelled_ag_truepos_s [label='p = {p_prophrate}'] transplant_notcancelled_ag_truepos_s-> proph_no_transplant_notcancelled_ag_truepos_s [label='p = {p_noprophrate}'] donor_neg_s -> ag_trueneg_s [label='p = {p_specificity}'] donor_neg_s -> ag_falsepos_s [label='p = {p_falsepositive}'] ag_falsepos_s -> transplant_cancelled_ag_falsepos_s [label='p = {p_cancelled}'] ag_falsepos_s -> transplant_notcancelled_ag_falsepos_s [label='p = {p_nocancelled}'] transplant_notcancelled_ag_falsepos_s -> proph_yes_transplant_notcancelled_ag_falsepos_s [label='p = {p_prophrate}'] transplant_notcancelled_ag_falsepos_s -> proph_no_transplant_notcancelled_ag_falsepos_s [label='p = {p_noprophrate}'] proph_yes_transplant_notcancelled_ag_truepos_s-> inf_y_proph_yes_transplant_notcancelled_ag_truepos_s [label='p = {p_breakthrough_donorpos}'] proph_yes_transplant_notcancelled_ag_truepos_s-> inf_n_proph_yes_transplant_notcancelled_ag_truepos_s [label='p = {p_nobreakthrough_donorpos}'] proph_no_transplant_notcancelled_ag_truepos_s-> inf_y_proph_no_transplant_notcancelled_ag_truepos_s [label='p = {p_transmission}'] proph_no_transplant_notcancelled_ag_truepos_s-> inf_n_proph_no_transplant_notcancelled_ag_truepos_s [label='p = {p_nontransmission}'] ag_falseneg_s->inf_y_ag_falseneg_s [label='p = {p_transmission}'] ag_falseneg_s->inf_n_ag_falseneg_s [label='p = {p_nontransmission}'] ag_trueneg_s->inf_y_ag_trueneg_s [label='p = {p_spont_cryptococcus}'] ag_trueneg_s->inf_n_ag_trueneg_s [label='p = {p_nospont_cryptococcus}'] proph_yes_transplant_notcancelled_ag_falsepos_s-> inf_y_proph_yes_transplant_notcancelled_ag_falsepos_s [label='p = {p_breakthrough_donorneg}'] proph_yes_transplant_notcancelled_ag_falsepos_s-> inf_n_proph_yes_transplant_notcancelled_ag_falsepos_s [label='p = {p_nobreakthrough_donorneg}'] proph_no_transplant_notcancelled_ag_falsepos_s-> inf_y_proph_no_transplant_notcancelled_ag_falsepos_s [label='p = {p_spont_cryptococcus}'] proph_no_transplant_notcancelled_ag_falsepos_s-> inf_n_proph_no_transplant_notcancelled_ag_falsepos_s [label='p = {p_nospont_cryptococcus}'] }} ") #Add grviz object to return_list return_list$plot<-grViz(grviz_text) #Final return return(return_list) } add_svg_annotation <- function(svg, lines, x = 40, y = 1200, fontsize = 50, lineheight = 1.3) { text_node <- xml_add_child( svg, "text", x = x, y = y, fill = "black", `font-size` = fontsize, `font-family` = "Helvetica" ) for (i in seq_along(lines)) { xml_add_child( text_node, "tspan", lines[[i]], x = x, dy = if (i == 1) "0" else as.character(fontsize * lineheight) ) } invisible(svg) } ``` ## Other results - [ **Additional tree analyses** ](additional_trees.qmd) : alternative clinical assumptions\- [ **One-way sensitivity analysis** ](tornado.qmd) : key drivers of results\- [ **Probabilistic sensitivity analyses** ](psa.qmd) : uncertainty and robustness\- [ **About** ](about.qmd) : methods, assumptions, and disclosures